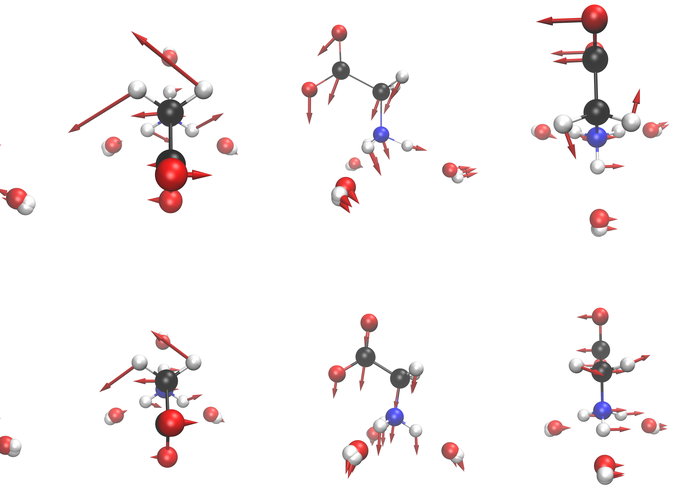
 Glycine Modes
Glycine Modes
 Glycine Modes
Glycine Modes
Proteins are biomolecules involved in cellular structure as well as function. These molecules are long chain polymers consisting of amino acids, which are organic compounds containing many different functional groups such as amine and carboxylic acids. Actin proteins form part of the cellular structure, membrane proteins act as channels for transfer of ions, and enzymes catalyze critical cellular reactions. While structure is a well-appreciated determinant of function for proteins, the roles of the dynamics of protein molecules and the surrounding solvent have been less well studied. In this talk, I will be presenting an analysis of the statistical fluctuations and dynamics of biomolecules and their interactions with solvent, starting from basic amino acids in water all the way up to the full complexity of enzymes, using molecular simulations. A combination of experimental and computational techniques is a powerful tool for obtaining insight into dynamical events in biomolecular systems. I have used force-field based classical molecular dynamics simulations using an advanced polarizable force field to study the behaviour of these biomolecules in solution and have simulated experimental observables to understand their various conformational motions. In particular, I am focussing on simulating the infra-red spectrum as well as the entropy of the biomolecules and the solvent as a function of their interactions, to understand the components and the effects of these molecular motions.